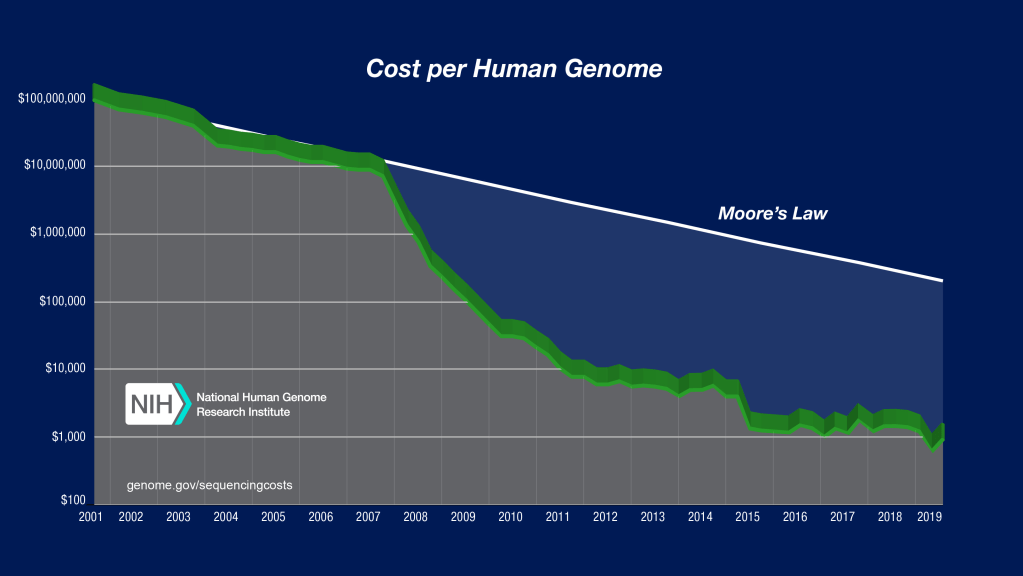
Genotyping technology has seen an outstandingly fast advancement in the last decades (Figure 1). In particular, after 2008, when “next-generation” sequencing appeared, sequencing improvement profoundly outpaced Moore’s law (genome.gov). As a consequence, the same amount of DNA that years ago required years to “read” and millions of Euros, today can be done in few hours and <1000 Euros.
Nevertheless, while brute genotyping power is a fundamental tool in any cutting-edge genotyping project, alone it is not sufficient (yet?) to achieve the aims of our project. Two main issues need to be addressed to obtain an enormous yet usable dataset of raw reads, and to be able to build a reliable reference genome from it: genome complexity and computational constraints (as said before, genotyping advancement outpaced Moore’s law). Fortunately, as we’ll see soon, the same approach can be the solution to both the problems.