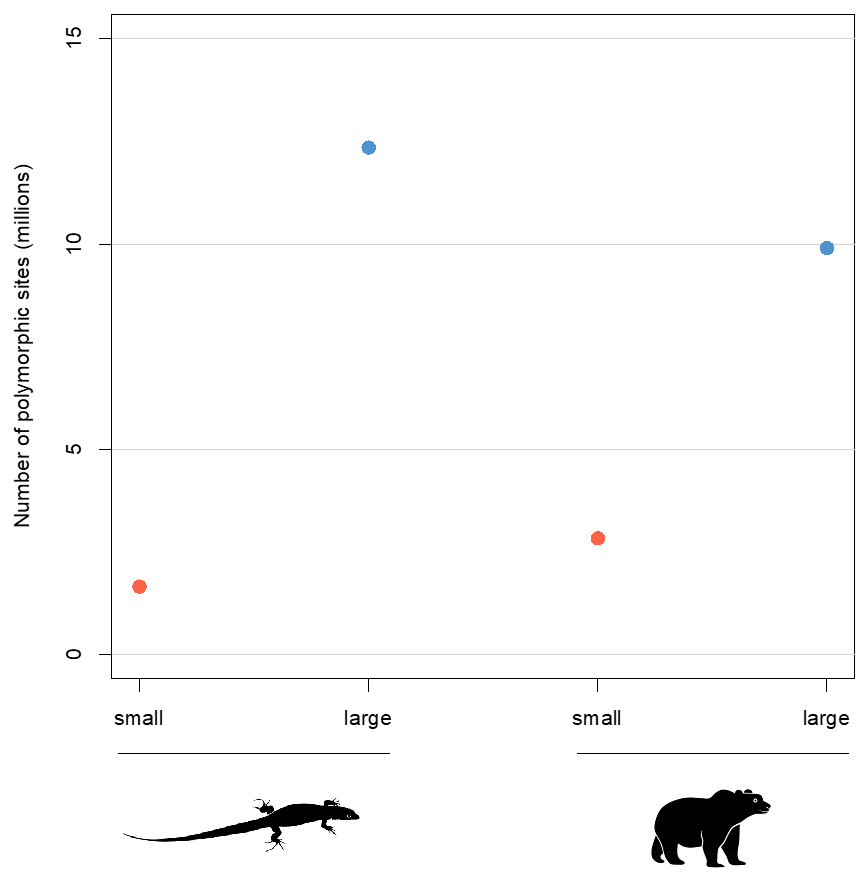
As soon as we received the resequencing data from individuals of the large and small population, respectively, for each of the five target species (details in the post: Here comes the data! 06.05.2021), we started processing and analyzing it to study genomic variation patterns and reconstruct demographic histories. These steps require a reference genome for each species that is used to map the resequencing data. In fact, this type of data represents small fragments of sequences that need to be recomposed into the individual’s genome, similarly to assembling a puzzle, and the reference genome provides the “image” to guide the appropriate placement (“mapping”) of these pieces. Considering that the specific reference genomes of our species are still in development, we started some preliminary analyses using closely related species with available reference genomes: Ursus maritimus (bear), Podarcis muralis (lizard), Bombina bombina (toad) and Acipenser ruthenus (sturgeon). After several trials, we excluded the butterfly from these preliminary analyses as no appropriately closely-related genomes are available so far and we preferred to wait for the specific reference, which will be ready soon. In the other four species, we started mapping our resequencing data to a reference, identifying genetic polymorphisms characterizing each sample, exploring genetic diversity within and between populations, inferring past demographic changes across time and estimating the burden of deleterious mutations (genetic load, see the post: Studying genomes to save endangered species 23.03.2021). Preliminary results in the bear and lizard suggest depleted genetic diversity in the small populations compared to the large ones (Figure below). We are looking forward to further investigating these valuable study systems and better understand their genomic susceptibility to extinction.